






 he combination of MR imaging with extrinsic contrast changing agents led many researchers to try to highlight anatomical and pathological structures and even metabolic processes which, per se, are invisible on single plain images [⇒ Rinck 1999].
he combination of MR imaging with extrinsic contrast changing agents led many researchers to try to highlight anatomical and pathological structures and even metabolic processes which, per se, are invisible on single plain images [⇒ Rinck 1999].

Image series of the same anatomical structure during a time period before and after the application of a contrast agent add another dimension, commonly described as perfusion imaging. Perfusion describes blood delivery to tissues, usually flow on the capillary level.
Perfusion imaging can be categorized into two classes: those methods monitoring target tissue signal changes after the application of an extrinsic contrast agent, and those relying upon intrinsic factors such as increased blood volume or blood oxygenation and flow in the micro-vasculature. The former can be visualized by dynamic imaging, the latter by functional imaging.
However, the individual images of a dynamic series will not always yield all information contained in them. Thus, additional processing has been suggested (Figure 16-01).
 This chapter gives an overview of some of the techniques. It cannot be exhaustive considering the wide spectrum of the field. For detailed scientific treatises see Maintz and Torheim [⇒ Maintz 1998, ⇒ Torheim 1999].
This chapter gives an overview of some of the techniques. It cannot be exhaustive considering the wide spectrum of the field. For detailed scientific treatises see Maintz and Torheim [⇒ Maintz 1998, ⇒ Torheim 1999].
Traditionally, the following methods have been applied to analyze dynamic contrast-enhanced images:
 visual inspection of the time-intensity curves;
visual inspection of the time-intensity curves;
 visual inspection of the time series by running the images in a movie;
visual inspection of the time series by running the images in a movie;
 superposition / subtraction images.
superposition / subtraction images.
Yet, the most intriguing and interesting way of processing the data is by the creation and subsequent visualization of parametric maps: images combining parametric images derived from the information present in the image series with anatomical information (Table 16-01).
To be clinically useful, such a postprocessing method must be robust, reliable as well as automatic or semi-automatic.
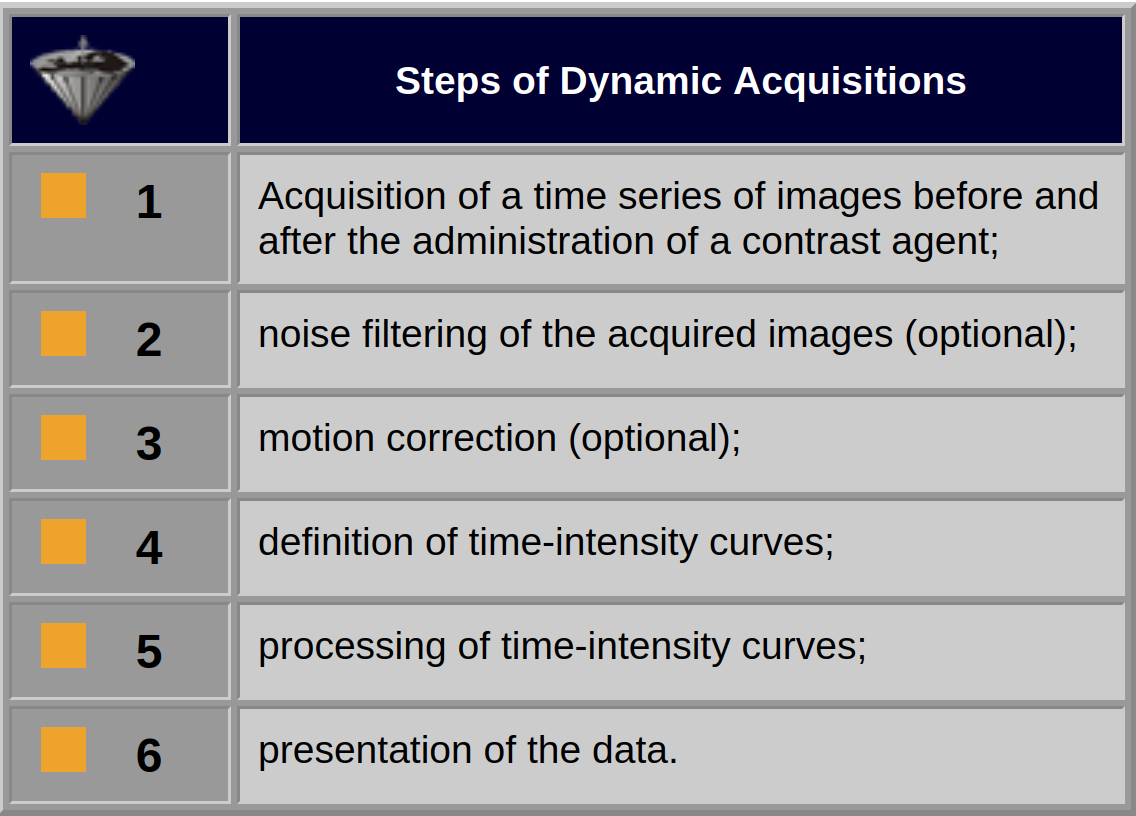
Table 16-01:
Steps of the entire dynamic acquisition and image-processing procedure. The noise-filtering and motion-correction steps are optional, but in many instances necessary and fairly complicated.